MCDIFFaX
DIFFaX is a software written by Mike Treacy and coworkers to calculate X-ray and neutron diffraction patterns of stacking disordered materials. There are many examples in the materials world - every layered material can in principle have some irregularities in how its layers are stacked. Famous examples include graphite, molybdenum sulphide, diamond and the MEL/MFI zeolitic system. Our work started off working on stacking disorder in ice and has since expanded to diamond, silver iodide, ammonium fluoride and many other materials.
There are many different ways in which stacking disordered ice (ice Isd) can be prepared in the lab. Depending on how ice Isd is made and its thermal history the stacking disorder will be different. This manifests mainly in different fractions of cubic and hexagonal stacking. The presence of memory effects within the stacking sequences can introduce additonal complexity.
To calculate a diffraction pattern, DIFFaX requires information about the atomic structure of the layers, the probabilities of the various types of stacking and the symmetry relationships between stacked layers. In addition, the calculated pattern is convolved with a peak-profile function.
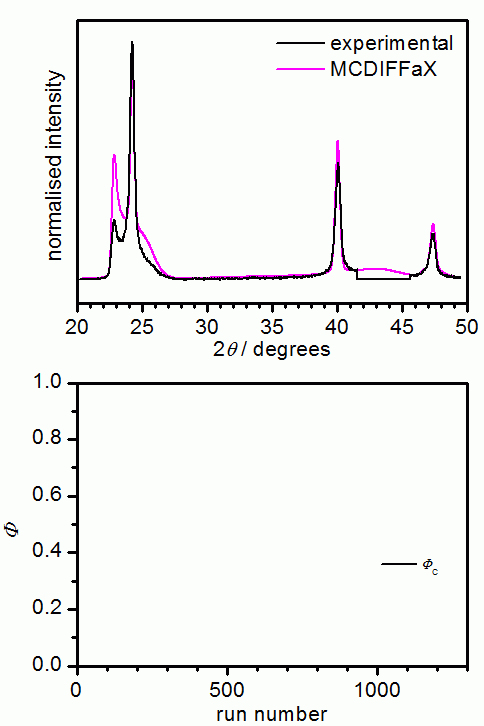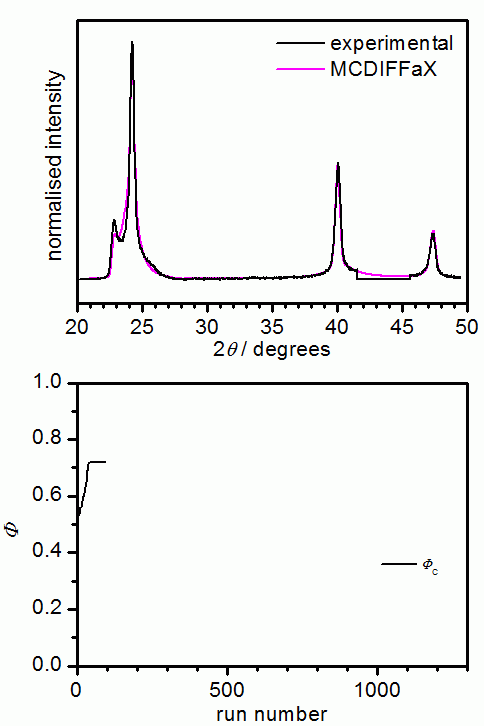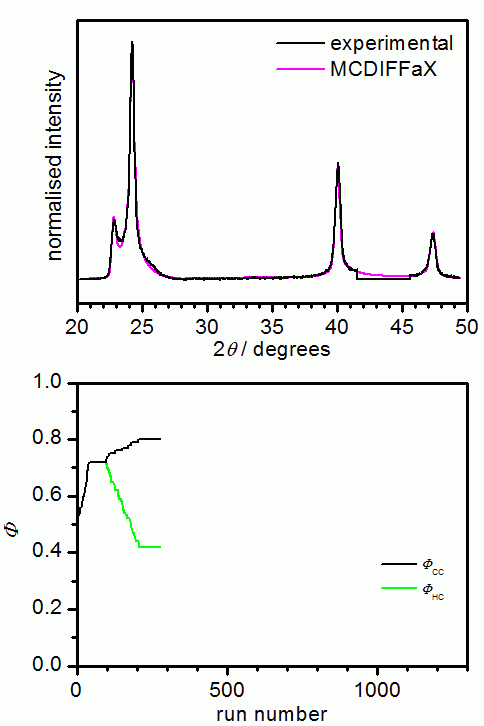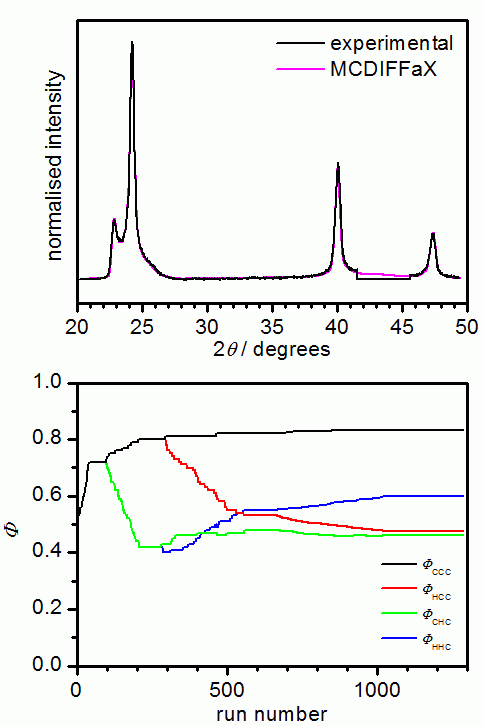
DIFFaX itself is not capable of refining the various parameters. This is why we have "embedded" DIFFaX in a least-square environment within our MCDIFFaX software. MCDIFFaX uses a simple Monte Carlo based approach to refine the various unknown parameters including stacking probabilities, lattice constants, displacement parameters, zero shift and peak-profile parameters (u, v, w and GL ratio). The programme randomly suggests a change in one of the parameters, DIFFaX is then asked to calculate the diffraction pattern and if this leads to an improvement of the fit, the change in the parameter is accepted. The idea is that this process goes on until the best possible fit to the experimental data is obtained. The user is required to specify which parameters should be refined and also the step size of a change. In addition, it is possible to allow a certain percentage of unfavourable changes in order to avoid ending up in a local minimum. Ideally, the step size is chosen quite large initially and then decreased successively as the fit converges. Similar to Rietveld refinements, MCDIFFaX requires the user to develop a bit of a "feeling" for the various parameters. For example, it is advisable to first refine the lattice constants and to then keep them constant while other parameters are optimised.
In our opinion, it is good practise to initially assume that there are no memory effects present in the sample. Once the best fit has been obtained, 1st, 2nd or even 3rd-order memory effects can be switched on in MCDIFFaX to investigate if this improves the fit to the pattern further.
LENCA (Local ENvironment ClAssification)
Using large-box structures of liquid mixtures from neutron diffraction studies, LENCA classifies local environments in binary mixtures as AA, AB or BB. This gives insights into the state of mixing. Our LENCA approach was used for a study into the structures of azeotropic mixtures, where either AB interactions dominate other interactions between the like molecules. LENCA offers an opportunity to break down complex interactions into readily understandable quantities.
RandomIce
Our RandomIce program produces large supercells of hydrogen-disordered ice structures. It requires a hydrogen-ordered structure as the initial input including the positions of unoccupied hydrogen sites. The program first builds a connectivity list of the hydrogen bonds. The disordering is achieved by using a “hopping” H+ throughout the structure. A hopping process carries on until the original starting point (OH-) is met. By defining probabilities for the jumping from different crystallographic sites, it is possible to prepare partially ordered hydrogen-disorderd structures in line with fractional occupancies determined from neutron diffraction. A full description of RandomIce is given here.
NanoVD
Our NanoVD program creates large supercell structures with defined numbers of Voronoi domains. The Voronoi domains are filled with crystal structures that have been rotated and shifted randomly including from stacking disordered materials. We used our NanoVD program to create nanocrystalline stacking disordered ices to explore if such structures are equivalent with low-density amorphous ice. Full details are given here.